Early Stage Investigators

Targeted mass spectrometry approaches to understand CART processing and receptor interactions
Dr. James Checco's project is a fundamental study examining the molecular-level processing and receptor interactions of cell-cell signaling peptides produced from the cocaine- and amphetamine-regulated transcript (CART). CART plays important roles in glucose regulation and incretin secretion and may serve as a novel target for several diseases, including type 2 diabetes.
Learn More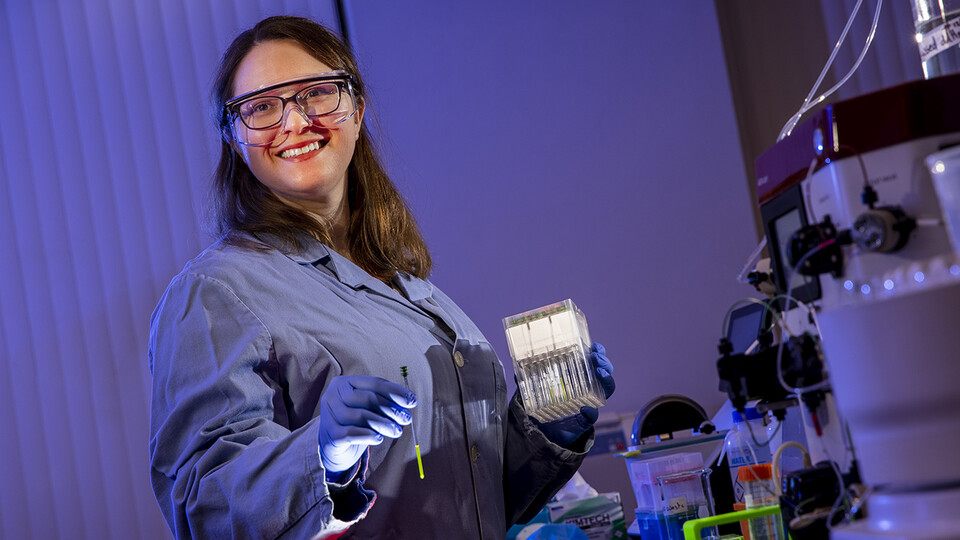
Investigating the structural basis for regulatory RNP assembly and function
Dr. Catherine Eichhorn's research uses integrated structural biology, biophysical, and chemical biology tools to investigate the folding and function of regulatory RNAs and RNPs. Her work aims to elucidate the biophysical properties governing RNP assembly and stability in order to achieve a rigorous understanding of how RNPs regulate gene expression in healthy individuals, and how dysfunction leads to divergent disease phenotypes.
Learn More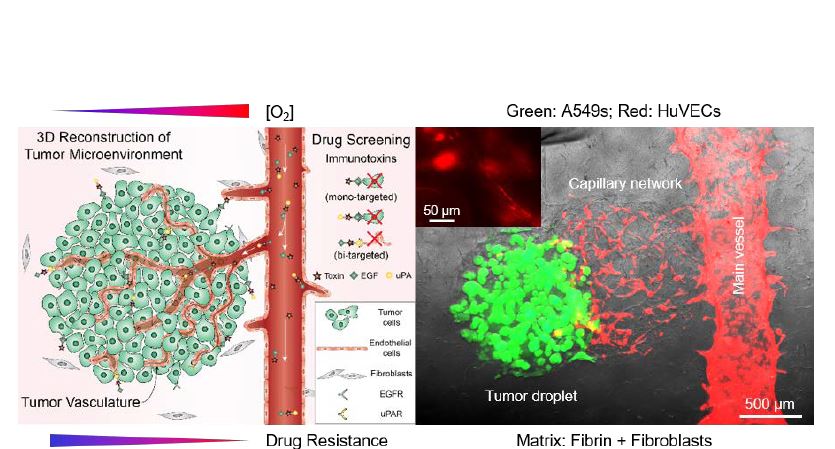
Investigating the structural basis for regulatory RNP assembly and function
Fanben Meng's overall research goal is to in vitro engineer biomimetic tumor constructs (Fig.1) via 3D bioprinting combinations of functional biomaterials and living cells, aiming to accurately model tumor complexity and heterogeneity. The main hypothesis will include: 1) 3D bioprinting can be used to precisely place ECMs and cells with the capability of dynamical tuning on bioink formulas; 2) Created heterogeneous microenvironments may promote cell-cell & cell-ECM interactions and therefore enhance the biological relevance of tissue constructs; and 3) Bioprinted models can serve as in vitro tools to predict cell response on therapeutic agents and treatments.Bioprinted models can serve as in vitro tools to predict cell response on therapeutic agents and treatments.
Learn More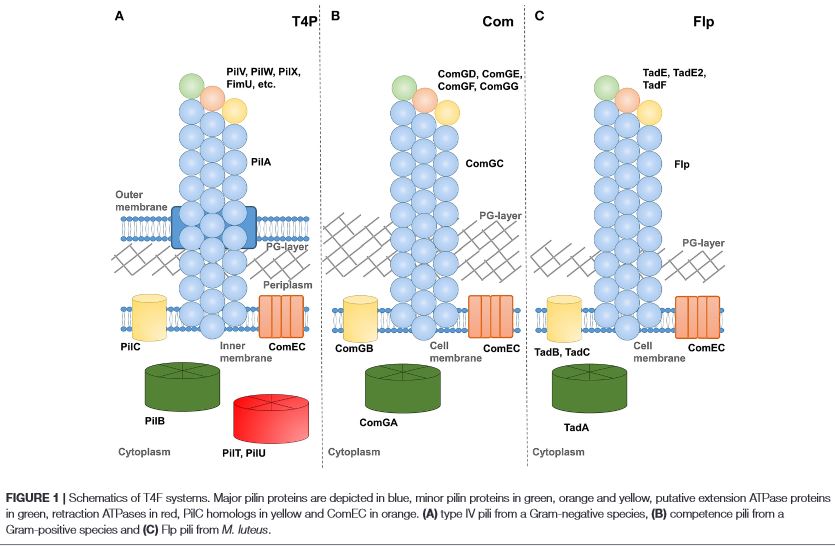
Molecular Mechanisms of Acinetobacter Motility and Adhesion
The goal of Dr. Piepenbrink’s research program is to elucidate the molecular mechanisms by which bacteria interact with their surroundings; this includes host cells, abiotic surfaces, extracellular structures and, in particular, other bacteria. Our group is approaching these basic questions by applying the lenses of structural biology and biophysics to microbial surface structures. These studies further seek to explain the phenotypic differences between bacterial strains of similar genetic background; typically, by comparing pathogenic bacteria to their commensal or environmental counterparts.
Learn More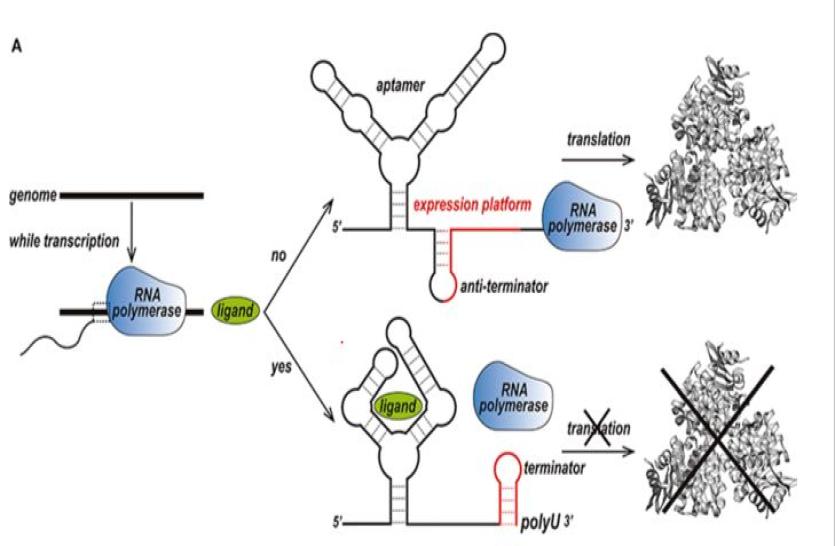
Towards a general predictive model of RNA tertiary structure energetics
Dr. Joseph Yesselmen's research on RNA nanotechnology offers exciting and potentially transformative applications for medicine. RNA can fold into complex tertiary structures that undergo conformational transitions in response to small-molecule binding. Proof-of-concept RNA machines that can respond to drug-like molecules have been engineered to control a wide range of biological processes. This ‘RNA chemostructural control’ offers exciting opportunities for the generation of ‘smart’ or user-controlled drugs on the horizon.
Learn MoreEstablished Investigators
Modeling and metabolic prediction of healthy and disease-promoting microbiomes
Led by Dr. Nicole Buan, this project is developing approaches to interpret and predict microbial phenotypes based on genomic information and environmental conditions in the host. By applying theoretical concepts onto empirical data, these approaches will be directly applicable to understanding host:microbe interactions that exacerbate or prevent diseases such as irritable bowel syndrome, Crohn’s Disease, and bacterial pathogenic infections.
Learn More
Hepatic Stabilin-2 Mechanisms of Clearance
The Harris lab is focused on receptor-mediated endocytosis in liver sinusoidal endothelial cells (SECs). SECs form the semi-permeable barrier between the blood and hepatocytes using fenestrated cytoplasm to allow molecules and not blood cells to perfuse through to the hepatocytes. SECs express several types of scavenger receptors, most notably the HARE or Stabilin-2 receptor which binds 14 known ligands. Several on-going projects in the lab address how the SECs respond to the accumulation of lipid droplets in the hepatocytes by way of non-alcoholic fatty liver disease (NAFLD) and the clearance rates of customized heparins by both Stabilin-1 and Stabilin-2.
Learn More
Micro-assembled Organoids-in-gels for High Throughput DrugScreening in Cancer Research
Organoids are three dimensional (3D) cell clusters that maintain the chemical, structural and organizational heterogeneity of tissues. They hold great potential as next-generation ex vivo disease models. Specifically, for cancer research, the characteristics of organoids and the advancement in methods for organoid generation make them ideal candidates for tumor models that recapitulate the inherent inter-/intra-patient heterogeneities associated with tumorigenesis and metastasis. Despite this promise, the current state of organoid culture is unable to generate large batches of organoids in microarray formats which are needed to facilitate discovery.
Learn More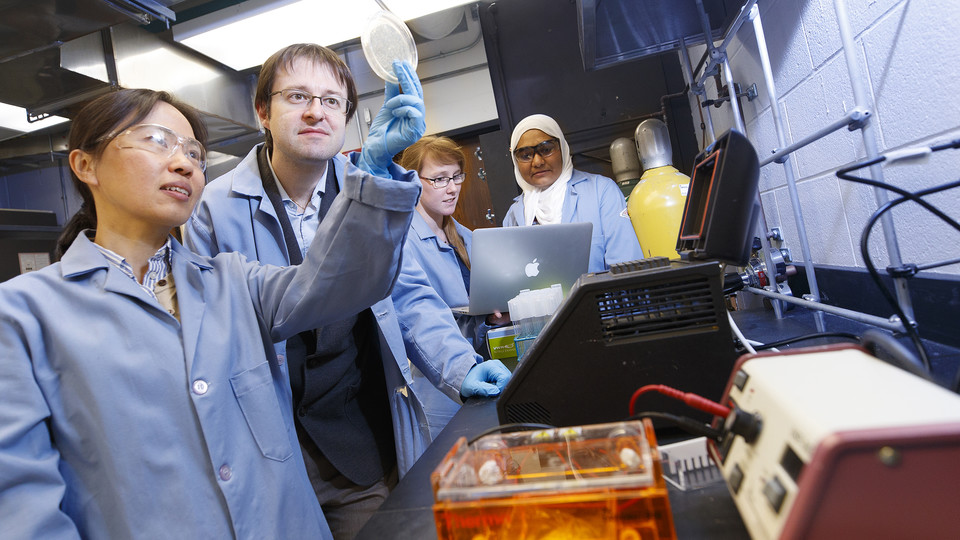
An Unnatural Amino Acid-based Chromatin Isolation Method (UChlMe) for the Study of Cancer-associated Protein-DNA Interactions
Protein-DNA interactions play key roles in many essential biological processes of all known organisms. Many human diseases, such as cancers, result from the misregulation of protein-DNA interactions. The long-term goal is to accurately map genome-wide and disease-associated protein-DNA interactions in single cells. Chromatin immunoprecipitation (ChlP) coupled to high-throughput sequencing (ChlP-Seq) has been widely used to map DNA sites of protein occupancy. Like most immunoprecipitation experiments, the specificity and affinity of antibodies is the most critical factor in ChlP analysis. It is estimated that fewer than 20% of tested antibodies are suitable for ChlP analyses. More importantly, it is still challenging to minimize random background noises from non-specific binding of antibodies. A novel method to elucidate genome-wide DNA binding sites of proteins with improved accuracy and sensitivity is critically needed.
Learn More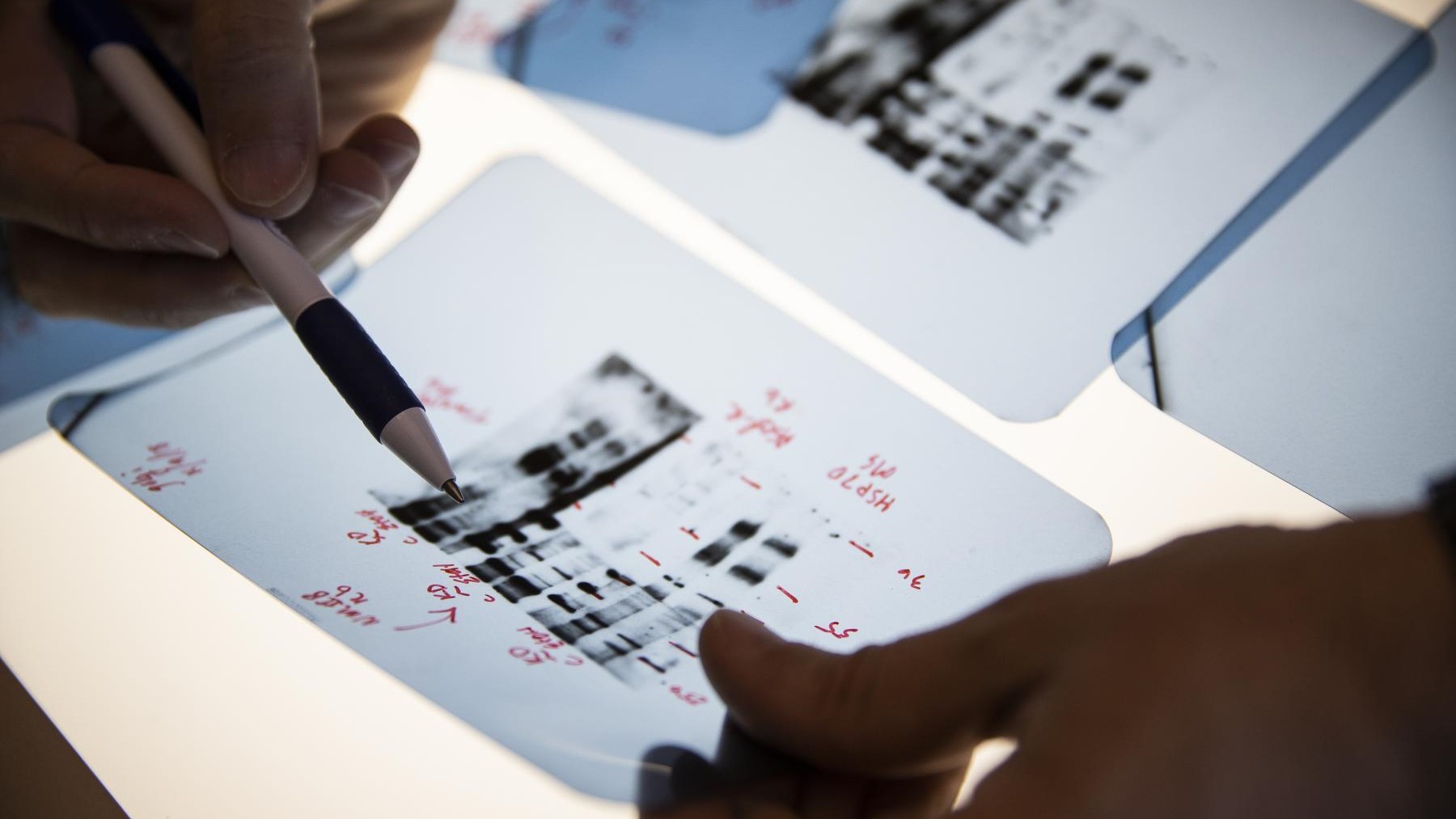
The role of N-glycan carrying Sialyl-Lewis (x) in progression of prostate cancer
Dr. Petrosyan's research project is entitled “The role for N-glycan carrying Sialyl-Lewis(x) in progression of prostate cancer”. Multiple studies have shown that serine protease Matriptase plays a substantial role in prostate cancer growth, invasion, and metastasis, but the mechanism of its activation at the cell surface is unknown. Matriptase undergoes glycan modification by the Golgi enzyme MGAT5, whose expression is also enhanced in advanced prostate cancer. Dr. Petrosyan hypothesizes that disorganization of the Golgi apparatus causes domination of MGAT5-mediated glycosylation resulting in the relocation of abnormally glycosylated Matriptase to the cell surface via its direct specific interaction with binding lectin, Galectin-3.
LEARN MORE
Intercellular Communication Between Epithelial Tissues Through Tight Junctions
Dr. Alex Vecchio's research aims to elucidate tight junction formation and deformation at the molecular level. These studies will clarify our understanding of tight junction structure−function and the biomolecular mechanisms that cause tight junction disruptions and lead to human disease.
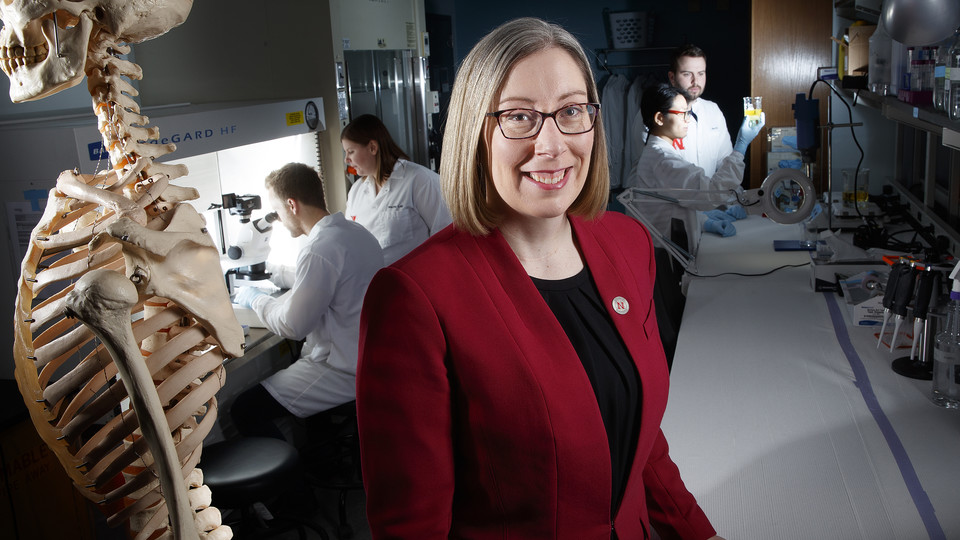

Identification and Biomaterial Targeting of Causative Biochemical Factors in Low Back Pain
The goals of Dr. Rebecca Wachs' work are to 1) understand the role of oxidative stress on development of chronic low back pain, and to 2) engineer targeted local pain therapeutics to alleviate opioid dependence. The majority of the population will experience low back pain in their lifetime, and many will go on to have chronicunresolved low back pain (LBP). Current conservative treatment for chronic LBP utilizes opioid prescriptions in ~30% of patients. LBP patients treated with opioids become addicted at a rate of 8-12%. An alternative approach that locally and directly address LBP could substantially decrease patient disability and reduce opioid dependence.
Learn MoreStress-Induced Pathway of Cell-Cell Adhesion Disruption in Pemphigus Pathogenesis
Dr. Ruiguo Yang's project aims to determine the role mechanical stress plays in regulating the pathogenesis of pemphigus, an autoimmune skin disease. With advanced single cell pair interrogation and stimulation techniques, the study will delineate the regulatory mechanisms in adhesive junctions during the cell-cell dissociation processes.
Learn More
A Multifunctional NO-Sensor Controlled by an Iron-Sulfur Cluster Switch
Dr. Limei Zhang's research addresses critical knowledge gaps in the mechanisms of survival and pathogenesis of Mycobacterium tuberculosis, the causative agent of tuberculosis which poses great threat to public health. Upon completion of the research, we expect to have a better understanding of how this pathogenic organism responds to nitric oxide stress, which is critical for its survival in the host. The outcomes are expected to offer new research directions for designing effective anti-tuberculosis therapeutics.
Learn More